Section: New Results
Computational Physiology
Computational Modeling of Radiofrequency Ablation for the Planning and Guidance of Abdominal Tumor Treatment
Participants : Chloé Audigier [Correspondant] , Hervé Delingette, Tommaso Mansi, Nicholas Ayache.
This work is carried out between the Asclepios research group, Inria Sophia Antipolis, France and the Medical Imaging Technologies, Healthcare Technology Center, Siemens Medical Solutions USA, Princeton, NJ.
Radio frequency ablation modeling, Patient-specific simulation, Lattice Boltzmann method, Computer model, Computational fluid dynamics, Heat transfer, Cellular necrosis, Parameter estimation, Therapy planning, Liver, Pre-clinical study, Medical imaging.
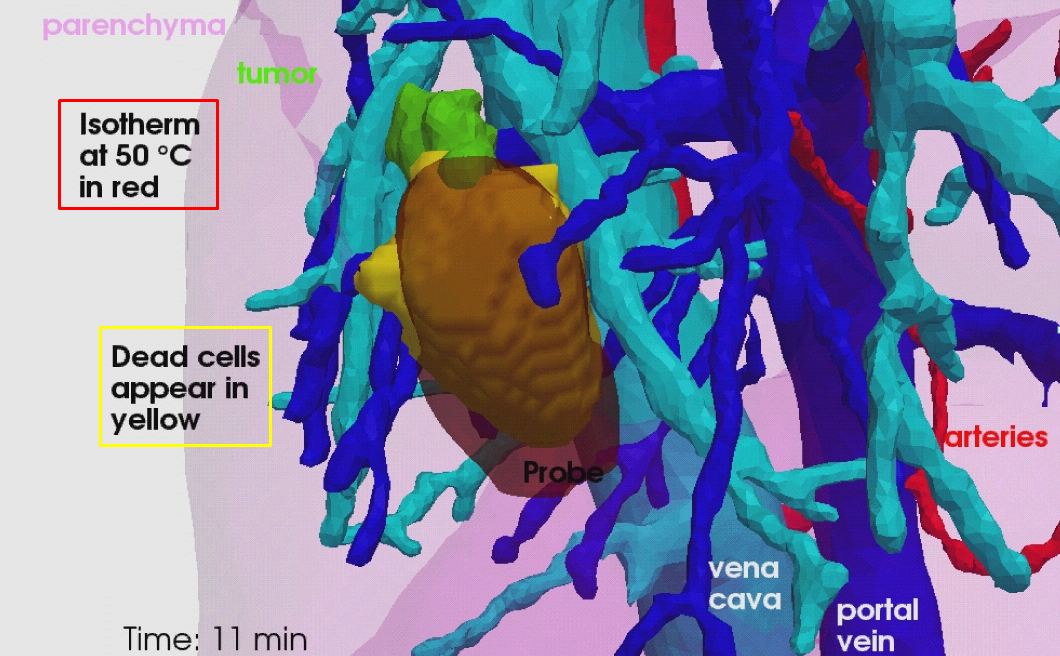
RFA is a minimally invasive therapy appropriated for liver tumor ablation. However, a patient-specific predictive tool to plan and guide the treatment is needed. We developed a computational framework for patient-specific planning of RFA with the following contributions:
-
A detailed computational model of the biophysical mechanisms (heat transfer, cellular necrosis, hepatic blood flow) involved in RFA of abdominal tumors based on patient images.
-
A new implementation of the bio-heat equations coupled with a cellular necrosis model using the Lattice Boltzmann Method (LBM) on Graphics Processing Units (GPU), which allows near real-time computation.
-
A CFD and porous media solver using LBM algorithm to compute the patient-specific blood flow in the hepatic circulatory system and the blood flow distribution inside the parenchyma.
-
A complete patient-specific geometry including hepatic venous and arterial circulation system.
-
The automatic estimation of the main parameters of the model. Two personalization strategies tested and evaluated on clinical and pre-clinical data.
-
The evaluation of the proposed model on a clinical dataset of ten patients.
-
The evaluation on a preclinical dataset of five swines from a comprehensive experimental set-up specially designed for RFA model validation.
Cardiac Electrophysiology Simulation for Arrhythmia Treatment Guidance
Participants : Rocio Cabrera Lozoya [Correspondant] , Maxime Sermesant, Nicholas Ayache.
Part of this work was funded by the European Research Council through the ERC Advanced Grant MedYMA 2011-291080 (on Biophysical Modeling and Analysis of Dynamic Medical Images).
Cardiac electrophysiology modeling, Intracardiac electrogram modeling, Radiofrequency ablation planning, Electroanatomical mapping.
-
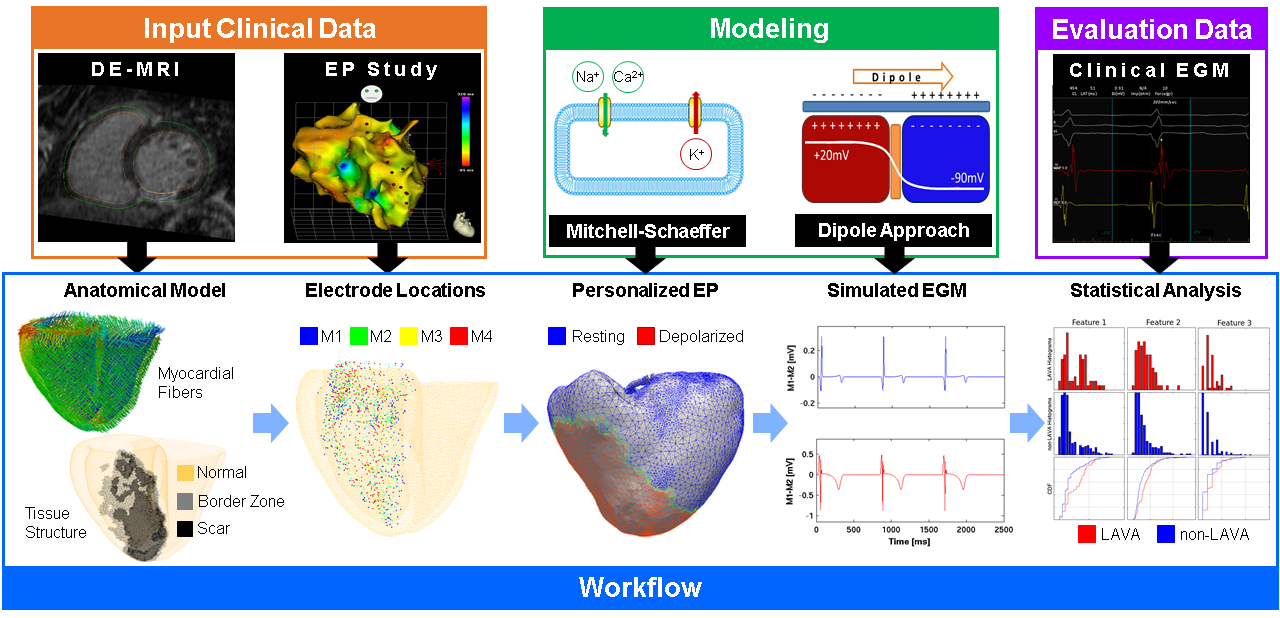
We developed silico patient-specific models constructed from 3D delayed-enhanced MRI to simulate intracardiac electrograms (EGM), including abnormal EGM as they are potential radiofrequency ablation targets (see Fig. 11) [14].
-
We derived a cardiac model using personalized electro-anatomical parameters and imaging data to define the underlying ventricular tachycardia (VT) substrate and predict re-entrant VT circuits [16].
Non-Invasive Personalisation of a Cardiac Electrophysiology Model from Body Surface Potential Mapping
Participants : Sophie Giffard Roisin [Correspondant] , Maxime Sermesant, Nicholas Ayache, Hervé Delingette.
This work has been supported by the European Project FP7 under grant agreement VP2HF (no 611823) and the ERC Advanced Grant MedYMA (on Biophysical Modeling and Analysis of Dynamic Medical Images).
Cardiac modeling, Personalised simulation, Inverse problem of ECG, Electrical simulation.
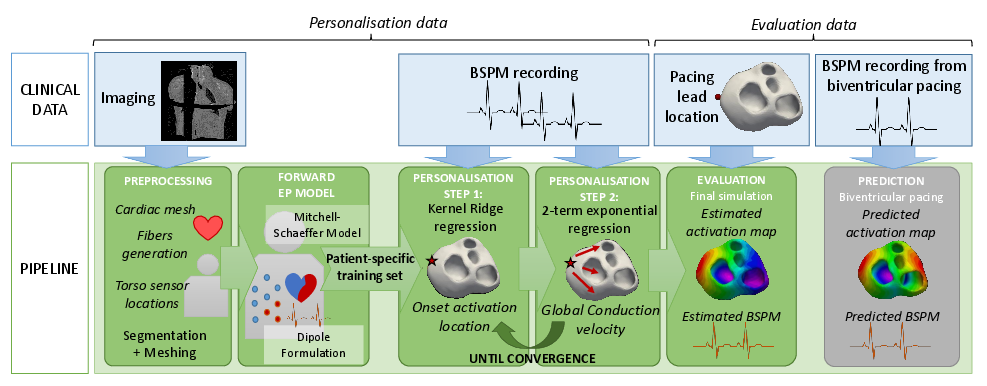
Within the VP2HF project, non-invasive cardiac electrical data has been acquired at the St Thomas' Hospital, London. It consists of Body Surface Potential Mapping (BSPM), which are recordings of the electrical potential on several locations on the surface of the torso. In [19], we use non-invasive data (BSPM) to personalise the main parameters of a cardiac electrophysiological (EP) model for predicting different pacing conditions (see Fig. 12). This is an encouraging first step towards a pre-operative prediction of different pacing conditions to assist clinicians for CRT decision and procedure. We have also worked on ECG data that are more commonly used in practice. In [38], we estimated the purkinje activation from 12-lead ECG using an intermittent left bundle brand block patient dataset.
Biophysical Modeling and Simulation of Longitudinal Brain MRIs with Atrophy in Alzheimer's Disease
Participants : Bishesh Khanal [Correspondant] , Nicholas Ayache, Xavier Pennec.
This work has been partly supported by the European Research Council through the ERC Advanced Grant MedYMA (on Biophysical Modeling and Analysis of Dynamic Medical Images).
Alzheimer's Disease (AD), Modeling brain deformation, Biophysical model, Simulation.
-
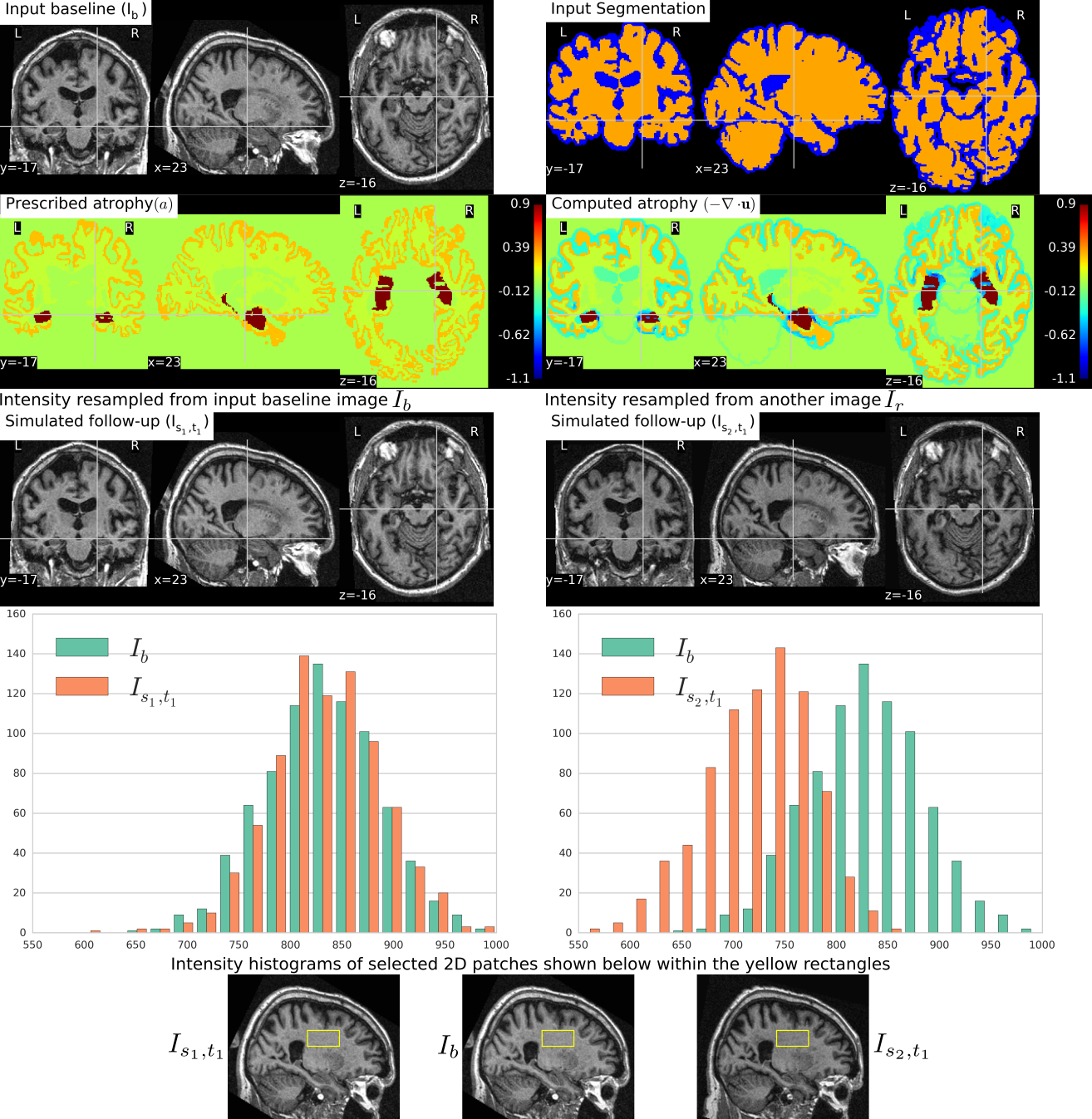
We completed a simulation tool that can simulate large databases of virtual realistic longitudinal MRIs with known volume changes[51]. This was based on our biophysical model of brain deformation due to atrophy in Alzheimer's Disease (AD)[25].
-
We have released our simulation software, named simul@trophy, as an open source software https://inria-asclepios.github.io/simul-atrophy/.
|
Brain Tumor Growth Personalization and Segmentation Uncertainty
Participants : Matthieu Lê [Correspondant] , Hervé Delingette, Jan Unkelbach, Nicholas Ayache.
This work is carried out between the Ascelpios research group, Inria Sophia Antipolis, France and the Department of Radiation Oncology of the Massachusetts General Hospital, Boston, USA.
Tumor growth, Radiotherapy, Modeling, Personalization, Segmentation, Uncertainty, Bayesian.
-
We elaborated a method for the synthesis of magnetic resonance images (MRIs) presenting glioblastoma [17].
-
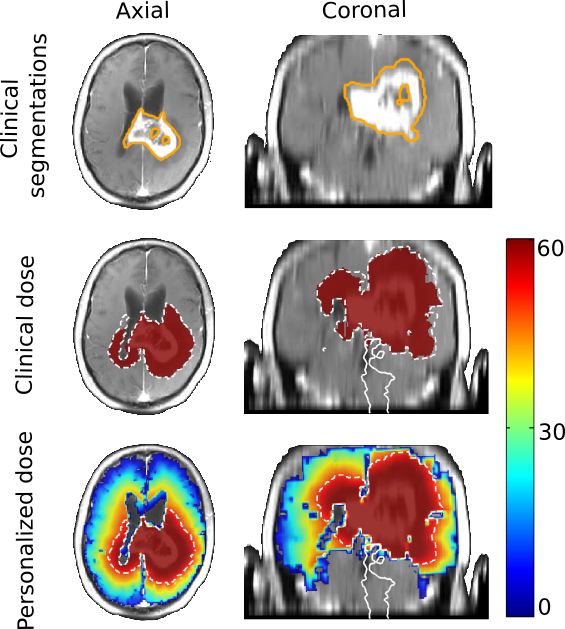
We elaborated a method for the sampling of several plausible segmentations, based on a single clinical one. This allows the uncertainty quantification of the radiotherapy plan based on several sample clinical target volumes [30].
-
We elaborated a method for the Bayesian personalization of a brain tumor growth model based on clinical MRIs [28].
-
We combined the segmentation sampling method with the tumor growth model personalization to personalize radiotherapy planning (see Fig. 14).
|
A Multiscale Cardiac Model for Fast Personalisation and Exploitation
Participants : Roch Philippe Molléro [Correspondant] , Xavier Pennec, Hervé Delingette, Nicholas Ayache, Maxime Sermesant.
This work has been partially funded by the EU FP7-funded project MD-Paedigree (Grant Agreement 600932) and contributes to the objectives of the ERC advanced grant MedYMA (2011-291080).
Cardiac modeling, Reduced model, Multi-fidelity modeling, Parameter estimation, Finite element mechanical modeling.
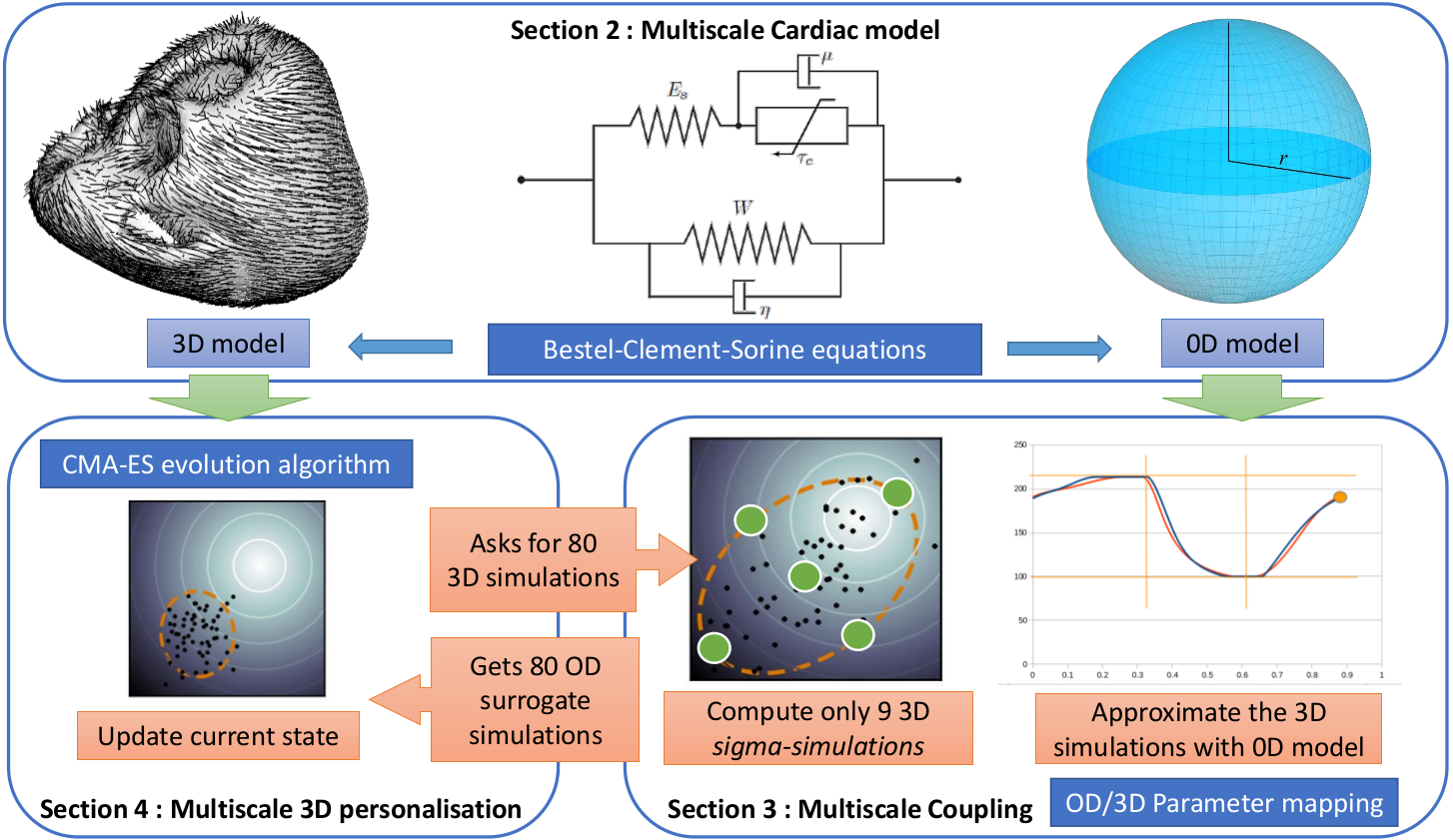
We developped a multi-fidelity 0D/3D cardiac model that allows us to get reliable (and extremely fast) approximations of the global behaviour of the 3D model with 0D simulations.
By making geometrical assumptions of symmetry, we first built a reduced 0D model of the heart which is very fast (15 beats/seconds). Then, we developped an original coupling method between the parameters of the 3D model and those of the 0D model. We used this multi-fidelity of the heart (in 0D and 3D) to speed-up an efficient optimization algorithm (the genetic algorithm CMA-ES) for the 3D model. As a result, we now have a fast personalisation method for the 3D model (see 15).
This methodology lead to a publication and poster presentation at the MICCAI Conference 2016 [41].
We applied this methodology in particuar to the cohort of 34 different heart geometries and data from the project MD-PAEDIGREE.